| HDAC4 |
|---|
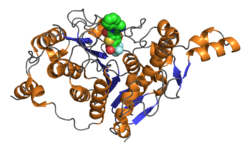 |
| Domaine catalytique Hdac4. PDB2vqj |
| Structures disponibles |
|---|
| PDB | Recherche d'orthologue: PDBe RCSB |
|---|
| Identifiants PDB |
|---|
2H8N, 2O94, 2VQJ, 2VQM, 2VQO, 2VQQ, 2VQV, 2VQW, 3UXG, 3UZD, 3V31, 4CBT, 4CBY, 5A2S |
|
|
| Identifiants |
|---|
| Aliases | HDAC4 |
|---|
| IDs externes | OMIM: 605314 MGI: 3036234 HomoloGene: 55946 GeneCards: HDAC4 |
|---|
| Position du gène (Homme) |
|---|
 | | Chr. | Chromosome 2 humain[1] |
|---|
| | Locus | 2q37.3 | Début | 239,048,168 bp[1] |
|---|
| Fin | 239,401,654 bp[1] |
|---|
|
| Position du gène (Souris) |
|---|
 | | Chr. | Chromosome 1 (souris)[2] |
|---|
| | Locus | 1|1 D | Début | 91,856,501 bp[2] |
|---|
| Fin | 92,123,421 bp[2] |
|---|
|
| Expression génétique |
|---|
| Bgee | | Humain | Souris (orthologue) |
|---|
| Fortement exprimé dans | - nerf sural
- muscle glutéal
- muscle gastrocnémien
- muscle tibial antérieur
- muscle de la cuisse
- muscle layer of sigmoid colon
- cellule endothéliale
- muscle deltoïde
- Skeletal muscle tissue of biceps brachii
- sang
|
| | Fortement exprimé dans | - otic vesicle
- granulocyte
- gastrula
- Rostral migratory stream
- dentate gyrus of hippocampal formation granule cell
- muscle de la cuisse
- lèvre
- septum interventriculaire
- tissu musculaire
- skeletal muscle tissue
|
| | Plus de données d'expression de référence |
|
|---|
| BioGPS |  | | Plus de données d'expression de référence |
|
|---|
|
| Gene Ontology |
|---|
| Fonction moléculaire | - NAD-dependent histone deacetylase activity (H3-K14 specific)
- sequence-specific DNA binding
- liaison ADN
- transcription corepressor activity
- histone deacetylase activity
- zinc ion binding
- transcription factor binding
- liaison d'histone déacétylase
- potassium ion binding
- liaison de chromatine
- liaison ion métal
- liaison protéique
- protein kinase binding
- activité hydrolase
- protein deacetylase activity
- identical protein binding
- promoter-specific chromatin binding
- RNA polymerase II cis-regulatory region sequence-specific DNA binding
- SUMO transferase activity
| | Composant cellulaire | - cytoplasme
- histone deacetylase complex
- Plaque motrice
- transcription repressor complex
- Sarcomère
- Z disc
- A band
- actomyosin
- noyau
- cytosol
- nucléoplasme
- complexe macromoléculaire
| | Processus biologique | - positive regulation of reactive oxygen species biosynthetic process
- histone H3 deacetylation
- skeletal system development
- Remodelage de la chromatine
- peptidyl-lysine deacetylation
- regulation of cardiac muscle contraction by calcium ion signaling
- negative regulation of glycolytic process
- response to interleukin-1
- regulation of transcription, DNA-templated
- regulation of gene expression, epigenetic
- cellular response to tumor necrosis factor
- positive regulation of DNA-binding transcription factor activity
- negative regulation of transcription by RNA polymerase II
- positive regulation of protein sumoylation
- negative regulation of DNA-binding transcription factor activity
- negative regulation of osteoblast differentiation
- transcription, DNA-templated
- neurodéveloppement
- regulation of striated muscle cell differentiation
- régulation positive de la transcription dépendante de l'ADN
- positive regulation of smooth muscle cell migration
- B cell activation
- histone H4 deacetylation
- cellular response to mechanical stimulus
- positive regulation of neuron apoptotic process
- cardiac muscle hypertrophy in response to stress
- positive regulation of cell population proliferation
- negative regulation of myotube differentiation
- response to denervation involved in regulation of muscle adaptation
- regulation of skeletal muscle fiber development
- osteoblast development
- B cell differentiation
- cellular response to parathyroid hormone stimulus
- réponse inflammatoire
- positive regulation of lamellipodium assembly
- regulation of protein binding
- negative regulation of transcription, DNA-templated
- histone deacetylation
- positive regulation of transcription by RNA polymerase II
- negative regulation of cell population proliferation
- positive regulation of smooth muscle cell proliferation
- organisation de la chromatine
- negative regulation of pri-miRNA transcription by RNA polymerase II
- protein deacetylation
- Sumoylation
- positive regulation of male mating behavior
| | Sources:Amigo / QuickGO |
|
| Orthologues |
|---|
| Espèces | Homme | Souris |
|---|
| Entrez | | |
|---|
| Ensembl | | |
|---|
| UniProt | | |
|---|
| RefSeq (mRNA) | |
|---|
NM_006037
NM_001378414
NM_001378415
NM_001378416
NM_001378417 |
| |
|---|
| RefSeq (protéine) | |
|---|
NP_006028
NP_001365343
NP_001365344
NP_001365345
NP_001365346 |
| |
|---|
| Localisation (UCSC) | Chr 2: 239.05 – 239.4 Mb | Chr 1: 91.86 – 92.12 Mb |
|---|
| Publication PubMed | [3] | [4] |
|---|
|
| Wikidata |
| Voir/Editer Humain | Voir/Editer Souris |
|

 Portail de la biologie cellulaire et moléculaire
Portail de la biologie cellulaire et moléculaire 


















